K-Means Clustering
K-Means is one of the most popular unsupervised learning algorithms. It partitions data into k clusters where each point belongs to the cluster with the nearest centroid.
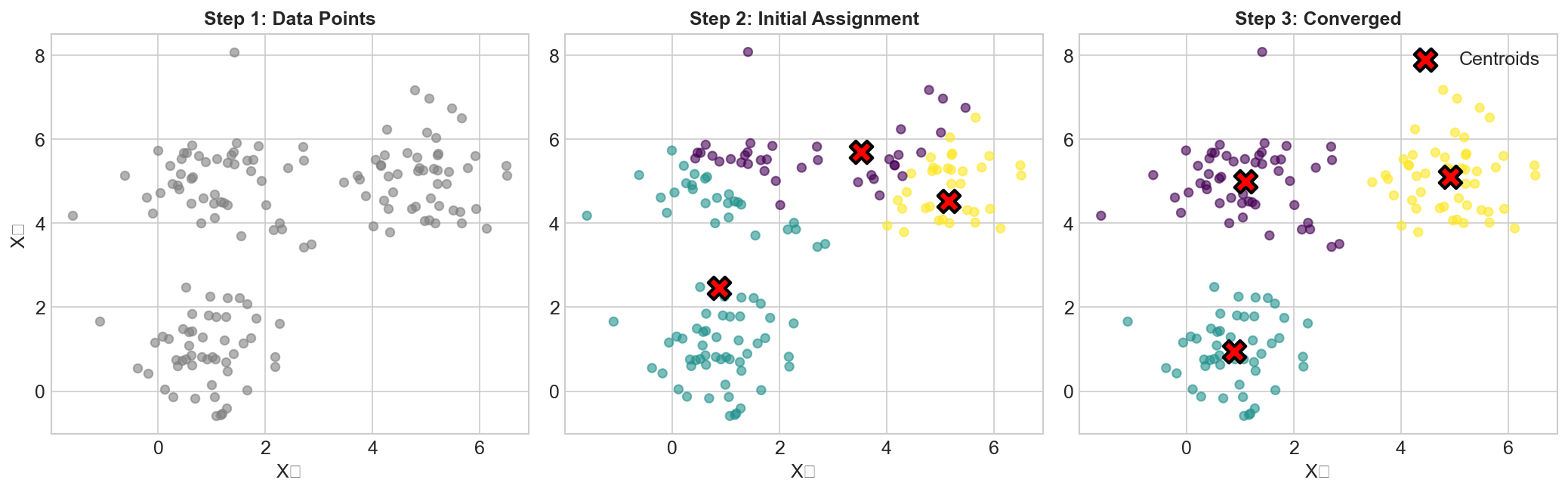
The Algorithm
Lloyd's Algorithm (Standard K-Means)
1. Initialize: Randomly place k centroids
2. Assign: Assign each point to nearest centroid
3. Update: Move centroids to mean of assigned points
4. Repeat 2-3 until convergence
Visual Example
Iteration 0: Iteration 1: Iteration 2:
× • • × • • × • •
• • × → • • × → • • ×
• • • • • • • • •
(converged)
× = centroid, • = data point
Mathematical Formulation
Minimize within-cluster sum of squares (WCSS):
J = Σᵢ Σₓ∈Cᵢ ||x - μᵢ||²
Where:
- Cᵢ is cluster i
- μᵢ is centroid of cluster i
Initialization Methods
Random
Randomly select k points as initial centroids.
- Simple but can lead to poor results
K-Means++
Smarter initialization:
- Choose first centroid randomly
- For each subsequent centroid:
- Compute distance to nearest existing centroid
- Choose new centroid with probability ∝ distance²
Much better results, default in sklearn.
Multiple Restarts
KMeans(n_clusters=5, n_init=10) # Run 10 times, keep best
Choosing K
Elbow Method
Plot WCSS vs k, look for "elbow":
WCSS
|\
| \
| \___
| \____
|___________
k
↑
elbow
Silhouette Score
Measures how similar points are to own cluster vs other clusters:
s = (b - a) / max(a, b)
a = mean distance to points in same cluster
b = mean distance to points in nearest other cluster
Range: [-1, 1], higher is better
Gap Statistic
Compare WCSS to expected WCSS under null reference distribution.
Advantages
- Simple and fast: O(nkt) where t is iterations
- Scales well: Works with large datasets
- Easy to interpret: Clear cluster assignments
- Guaranteed convergence: Always converges (to local minimum)
Limitations
- Must specify k: Number of clusters required upfront
- Sensitive to initialization: Can converge to local minima
- Assumes spherical clusters: Struggles with elongated shapes
- Sensitive to outliers: Mean is affected by outliers
- Equal-sized clusters: Tends to create similar-sized clusters
When K-Means Fails
✗ Non-spherical clusters (crescents, rings)
✗ Clusters of very different sizes
✗ Clusters of very different densities
✗ High-dimensional sparse data
Variants
K-Medoids (PAM)
Use actual data points as centers instead of means.
- More robust to outliers
- Works with any distance metric
Mini-Batch K-Means
Use random subsets for updates.
- Much faster for large datasets
- Slight quality tradeoff
K-Means with Cosine Similarity
Normalize vectors, use cosine distance.
- Better for text/sparse data
Bisecting K-Means
Recursively split clusters.
- More deterministic
- Can produce hierarchy
Practical Tips
Feature Scaling
from sklearn.preprocessing import StandardScaler
X_scaled = StandardScaler().fit_transform(X)
kmeans.fit(X_scaled)
Essential! K-means uses distances.
Dimensionality Reduction
from sklearn.decomposition import PCA
X_reduced = PCA(n_components=50).fit_transform(X)
kmeans.fit(X_reduced)
Helps with high-dimensional data.
Code Example
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
# Fit model
kmeans = KMeans(n_clusters=5, init='k-means++', n_init=10, random_state=42)
labels = kmeans.fit_predict(X_scaled)
# Evaluate
print(f"Inertia: {kmeans.inertia_}")
print(f"Silhouette: {silhouette_score(X_scaled, labels)}")
# Get centroids and assignments
centroids = kmeans.cluster_centers_
assignments = kmeans.labels_
Alternatives to Consider
| Algorithm | Best For |
|---|---|
| DBSCAN | Unknown k, arbitrary shapes, noise |
| Hierarchical | Need hierarchy, small data |
| Gaussian Mixture | Soft assignments, elliptical clusters |
| Spectral | Non-convex clusters |
| HDBSCAN | Varying densities, automatic k |
Key Takeaways
- K-Means partitions data into k clusters by minimizing within-cluster variance
- Use K-Means++ initialization (sklearn default)
- Always scale features before clustering
- Use elbow method or silhouette score to choose k
- Assumes spherical, equally-sized clusters
- Consider DBSCAN or GMM for non-spherical clusters